Two leading sequencing techniques are no longer at odds, thanks to an international effort led by scientists at University of California San Diego.
In a study published July 27, 2023 in Nature Biotechnology, the researchers debuted a new reference database called Greengenes2, which makes it possible to compare and combine microbiome data derived from either 16S ribosomal RNA gene amplicon (16S) or shotgun metagenomics sequencing techniques.
“This is a significant moment in microbiome research, as we’ve effectively rescued over a decade’s worth of 16S data that might have otherwise become obsolete in the modern world of shotgun sequencing,” said senior author Rob Knight, Ph.D., professor in the departments of Pediatrics at UC San Diego School of Medicine and Bioengineering and Computer Science at UC San Diego Jacobs School of Engineering. “Standardizing results across these two methods will significantly improve our chances of discovering microbiome biomarkers for health and disease.”
Microbiome studies depend on scientists’ ability to identify which microorganisms are present in a sample. To do this, they sequence the genetic information in the sample and compare it to reference databases that list which sequences belong to which organisms. 16S and shotgun sequencing are the two techniques most widely used in microbiome research, but they often yield different results.
“Many researchers assumed that data from 16S and shotgun sequencing were simply too different to ever be integrated,” said first author of the study Daniel McDonald, Ph.D., scientific director of The Microsetta Initiative at UC San Diego School of Medicine. “Here we show that is not the case, and provide a reference database that researchers can now use to do just that.”
The original Greengenes database had been widely used in the microbiome field for well over a decade. It was the reference database used by notable projects including the National Institutes of Health Human Microbiome Project, the American Gut Project, the Earth Microbiome Project and many others.
However, one of its fundamental limitations was that it relied on the sequence of a single gene, 16S, to identify the organisms in a sample. This well-studied gene has long been used as a taxonomic marker, with each organism having its own 16S “barcode.” This method can describe the contents of a microbiome sample with genus-level resolution, but it cannot always identify specific species or strains of microbes, which is important for clinical work.
Modern microbiome studies have since transitioned to using shotgun sequencing, which looks at DNA from all over the organisms’ genomes, rather than focusing on only one gene. This powerful approach gives researchers more species-level specificity and also provides insight into the microbes’ function.
Scientists often attributed the discrepancies between the two techniques to differences in the way the samples are prepared in the lab. However, the new study demonstrates that incompatibilities between the two techniques arise from differences in computation, where a better reference database allows for the same conclusions to be drawn from both methods. This addresses an important issue in the reproducibility of microbiome research and allows the re-use of data from millions of samples in older studies.
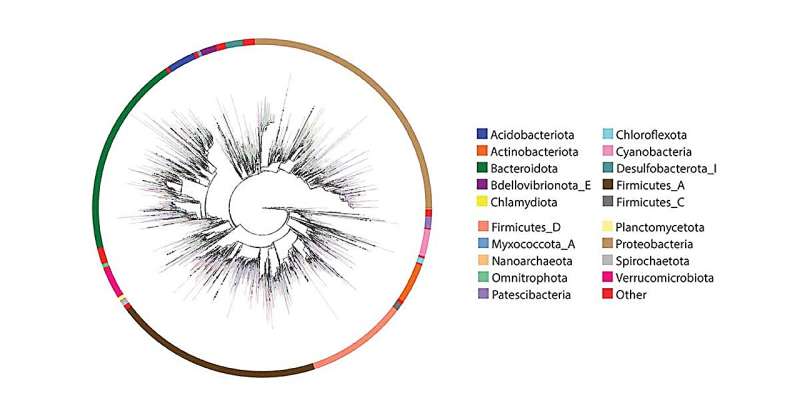
In trying to resolve these incompatibilities, the researchers first expanded the Web of Life whole genome database. They then used several new computational tools developed with co-author Siavash Mirarab, Ph.D., associate professor at UC San Diego Jacobs School of Engineering, to integrate existing high-quality full-length 16S sequences into the whole-genome phylogeny.
With another machine learning tool developed by Mirarab’s group, they placed 16S fragments from over 300,000 microbiome samples. The result was an expansive reference database that both 16S and shotgun sequencing data could be mapped onto.
To confirm whether Greengenes2 would help standardize findings from either sequencing technique, the researchers acquired both 16S and shotgun sequencing data from the same human microbiome samples and analyzed them both against the backdrop of the Greengenes2 phylogeny. The results from both techniques showed highly correlated diversity assessments, taxonomic profiles and effect sizes—something researchers had not seen before.
“Through Greengenes2, a huge repository of 16S data can now be brought back into the fold and even combined with modern shotgun data in new meta-analyses,” said McDonald. “This is a major step forward in improving the reproducibility of microbiome studies and strengthening physicians’ ability to draw clinical conclusions from microbiome data.”
More information:
Daniel McDonald et al, Nature Biotechnology (2023). www.nature.com/articles/s41587-023-01845-1
Provided by
University of California – San Diego
Citation:
New resource harmonizes 16S and shotgun sequencing data for microbiome research (2023, July 27)
retrieved 27 July 2023
from https://phys.org/news/2023-07-resource-harmonizes-16s-shotgun-sequencing.html
This document is subject to copyright. Apart from any fair dealing for the purpose of private study or research, no
part may be reproduced without the written permission. The content is provided for information purposes only.









